Analysis of nuclear morphology and its evolution
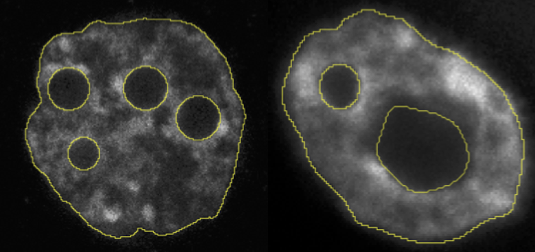
Nuclear membrane segmentation from DNA markers
The nuclear shape of mammals embryo is a major, yet unexplored, feature of embryogenesis. Studying its evolution requires an accurate segmentation of the nuclear membrane. To compensate for non-homogeneity of DNA fluorescent markers (e.g. DAPI staining), it is crucial to rely on shape regularized segmentation tools and region statistics.
Filamentous structure analysis
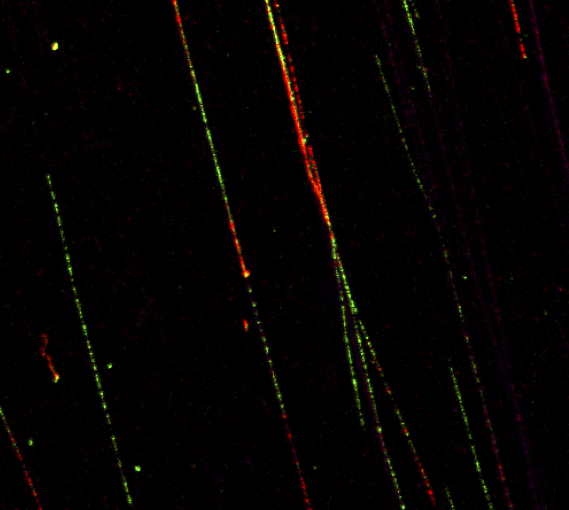
Automatic quantification of DNA fibers
The replication of DNA can be monitored at a single molecule level using the DNA fibers technique. By using on the one hand image processing techniques for detection of the fibers, and on the other hand classification techniques for labelling, it is possible to automatize such labor intensive task.
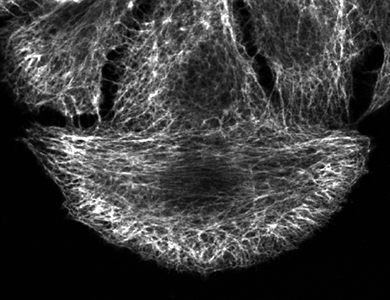
Analyzing the filaments structure: a spectral approach
The organisation of filament structures, such as the cytoskeleton of cells, is an important phenotypic feature describing the effects of biological materials on the cells (e.g. proteins or genes). Different filaments organization can be considered as different textures and the quantification ca be led by a spectral analysis.
See reference: Osmani 2018.
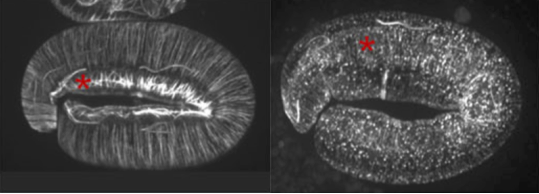
Analyzing filaments shapes: a geometrical approach
The geometry of single filaments, such as instance the epidermal microtubules of c. elegans worm, can be affected by the effects of biological material (e.g. proteins or genes). Geometrical properties of filaments can be used to evaluate any effect on the shape of filaments.
See reference: Quintin 2016.
Features organization analysis in mouse embryos
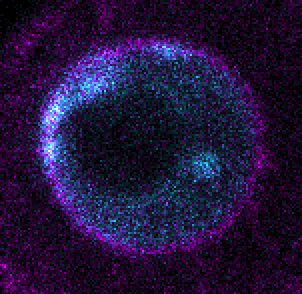
Localisation of DNA replication and RNA transcription
he precise location in the nuclei of DNA replication or RNA transcription for a particular gene is of great importance for understanding genomic events during embryogenesis. Such phenomena appear as spots within the three-dimensional space of the nuclei. Once the segmentation of the nuclear membrane and the detection of the spots are performed, many useful measures can be conducted (e.g. shortest distance to membrane).
See reference: Borsos 2019.
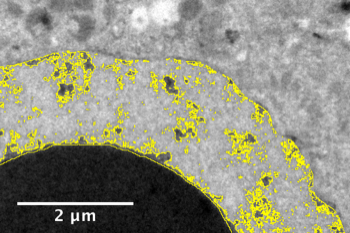

Heterochromatin quantification in mouse’s embryo with electron microscopy
The genetic material contained in cells is not only composed by genes, but also by a lot of other structures, such as heterochromatin. One way to quantify the compaction of heterochromatin is to quantify the proportion of electron-denses regions in electron microscopy images of cells.
See reference: Boskovic 2014.
Human fetal growth analysis
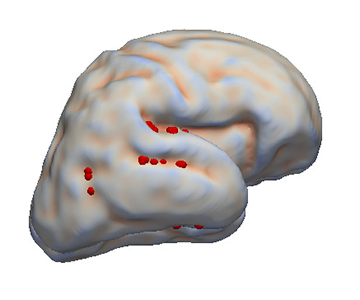
Feature selection for the study of the fetal brain folding
The fetal brain folding is an important process of the brain maturation. Taking advantage of the time-dependent shape information given by the atlas construction (see the atlas construction paragraph), the cortical plate can be reduced to a few cortical points describing the growth pattern.
See references: Pontabry 2017; Pontabry 2014; Pontabry 2013d; Pontabry 2013b.
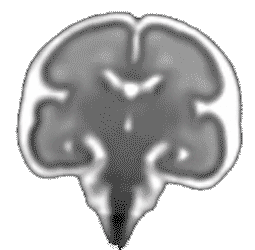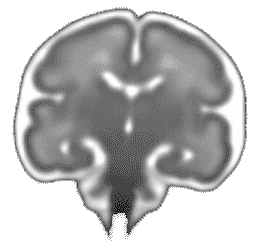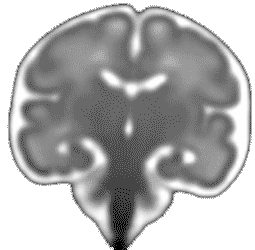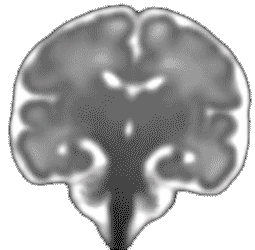
Atlas construction for the modeling of human brain maturation
A model of the human brain normal development is useful to study the development and to compare a new brain with the development trajectory. Such model can be constructed using only data (MRI images of brains) of a population with different ages and without forcing any prior information during its construction.
See references: Pontabry 2013d; Pontabry 2013c; Pontabry 2012.
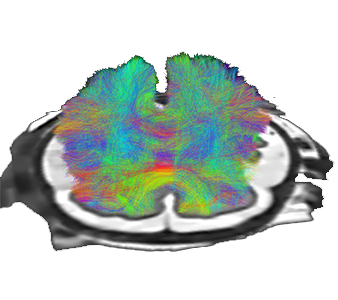
Tracography in diffusion weighted MRI
By using indirect observations (diffusion weighted MRI) of the neural fibers, we are able to reconstruct the full brain network. The uncertainty of such reconstruction is carried out here by a stochastic method (Markov chain Monte Carlo) that allow to process poor quality data, such as fetal brain MRI.
See references: Pontabry 2013d; Pontabry 2013a; Pontabry 2011; Rousseau 2011.